The Microbial World: Unseen Forces Shaping Our Lives
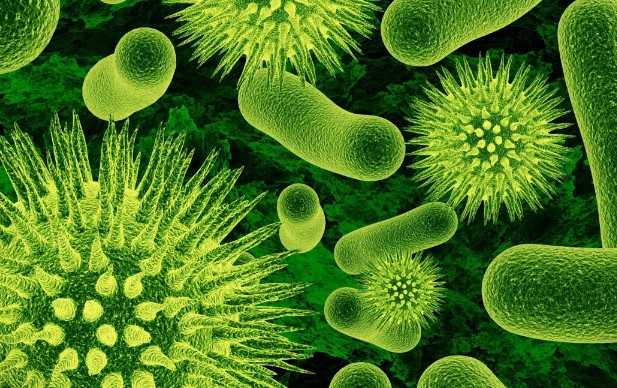
The Microbial Symphony: Unseen Forces Shaping Our Lives
Microbes: The Invisible Architects of Life
Microbes, though often overlooked due to their minuscule size, function as the unsung architects of life on Earth. Beyond aiding digestion and influencing climate patterns, these tiny organisms are integral to the very fabric of our existence. In the intricate dance of ecosystems, microbes contribute to nutrient cycling, soil fertility, and even the regulation of our own bodies. From the microscopic depths of our gut to the vast expanses of soil, the impact of microbes is profound, underscoring their role as the invisible thread connecting life forms and shaping the world as we know it.
The Hidden Complexity: Microbial Relationships and Environmental Dynamics
Delving into the microbial world reveals a hidden complexity that defies its size. Microbes engage in intricate relationships with each other and their environments, orchestrating a symphony of interactions. These relationships extend beyond simple processes, influencing the health of ecosystems and determining the success of various life forms. Understanding the dynamics of microbial communities opens doors to scientific breakthroughs, offering insights into how these unseen forces contribute to environmental resilience, biodiversity, and the delicate balance that sustains life.
Unveiling Potential: Microbial Applications in Medicine, Agriculture, and Conservation
As our knowledge of microbial ecosystems expands, so does our awareness of their potential applications. Microbes are not just players in the background; they hold the key to advancements in medicine, agriculture, and environmental conservation. From harnessing beneficial microbes for soil health and crop yield improvement to exploring the role of microbiota in personalized medicine, the microbial world is a treasure trove of possibilities. Embracing this microscopic frontier can lead to innovative solutions that address global challenges, demonstrating that even the tiniest entities can wield a monumental influence on the future of our planet.
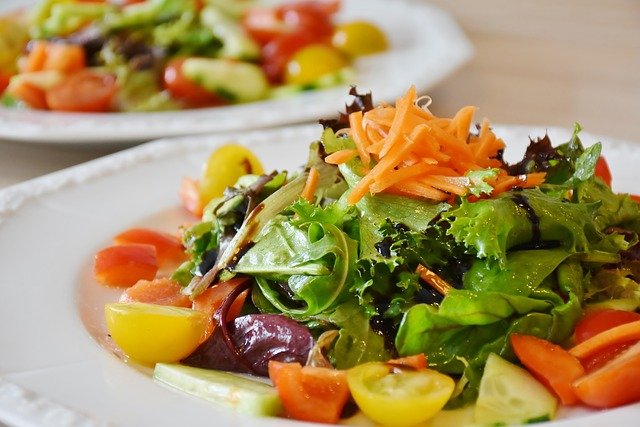











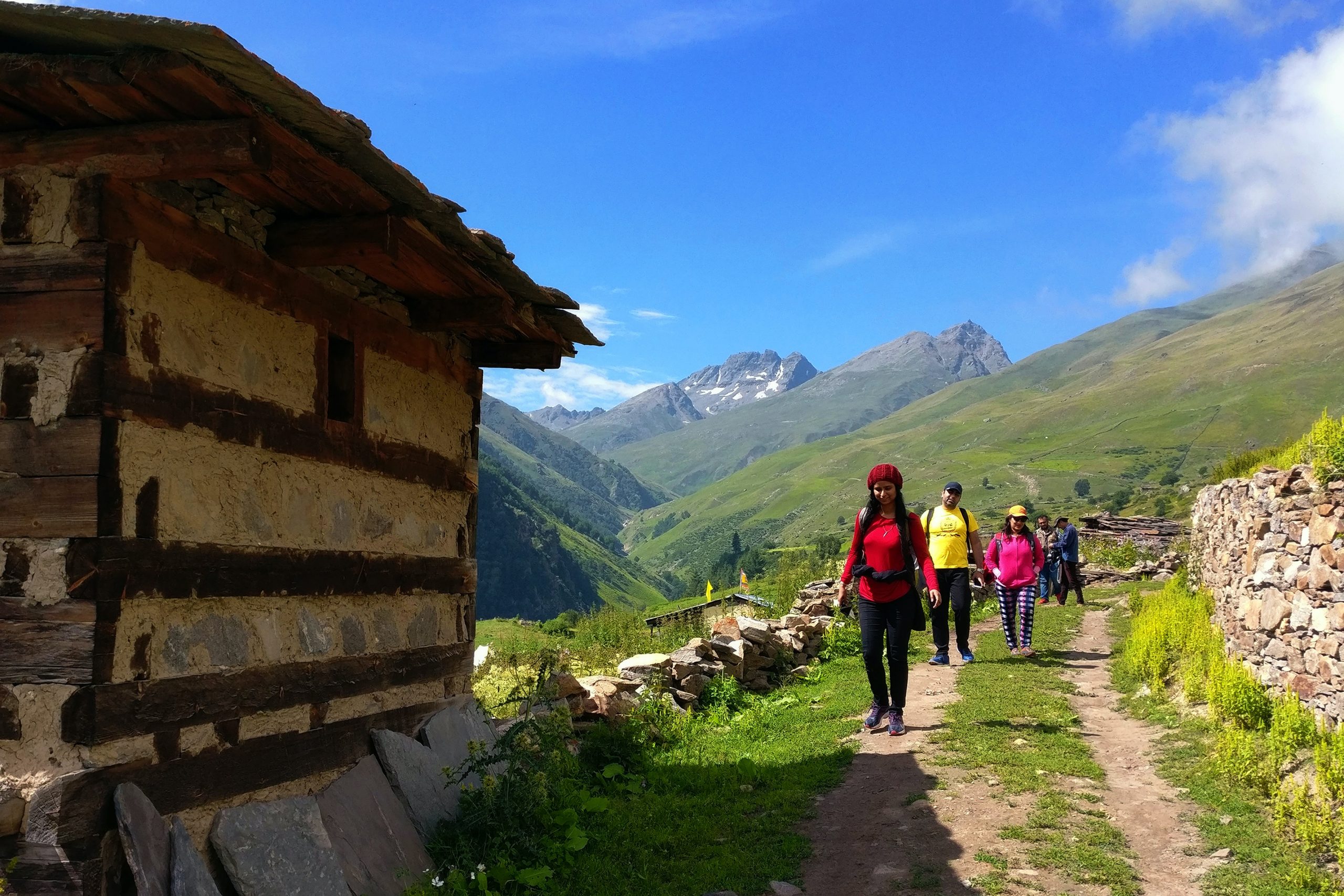
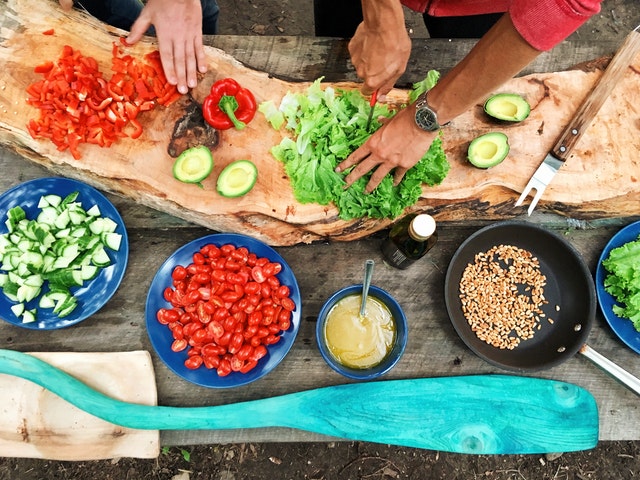
 Train journeys can be an exciting way to explore Germany, but it’s important to ensure you’re fueling your body with healthy food options. In this article, we’ll explore the importance of healthy eating on train journeys and what to look for when choosing healthy food options on German trains.
Train journeys can be an exciting way to explore Germany, but it’s important to ensure you’re fueling your body with healthy food options. In this article, we’ll explore the importance of healthy eating on train journeys and what to look for when choosing healthy food options on German trains.










 Losing your hair is frightening. However, for some, it is inevitable due to some illnesses such as
Losing your hair is frightening. However, for some, it is inevitable due to some illnesses such as 










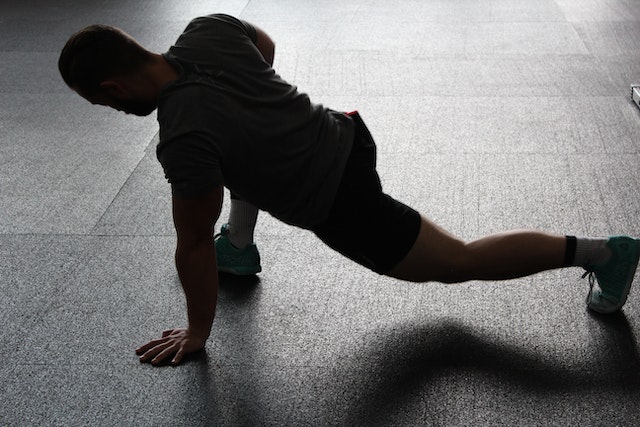
 In case you’ve got a passion for fitness and sports , then you need to make sure you know where the sporting goods stores are. Though you might not have a problem with shopping online, you might be passing up some of merchandise and the exclusive offerings that might not be listed online. If you want it to be
In case you’ve got a passion for fitness and sports , then you need to make sure you know where the sporting goods stores are. Though you might not have a problem with shopping online, you might be passing up some of merchandise and the exclusive offerings that might not be listed online. If you want it to be 










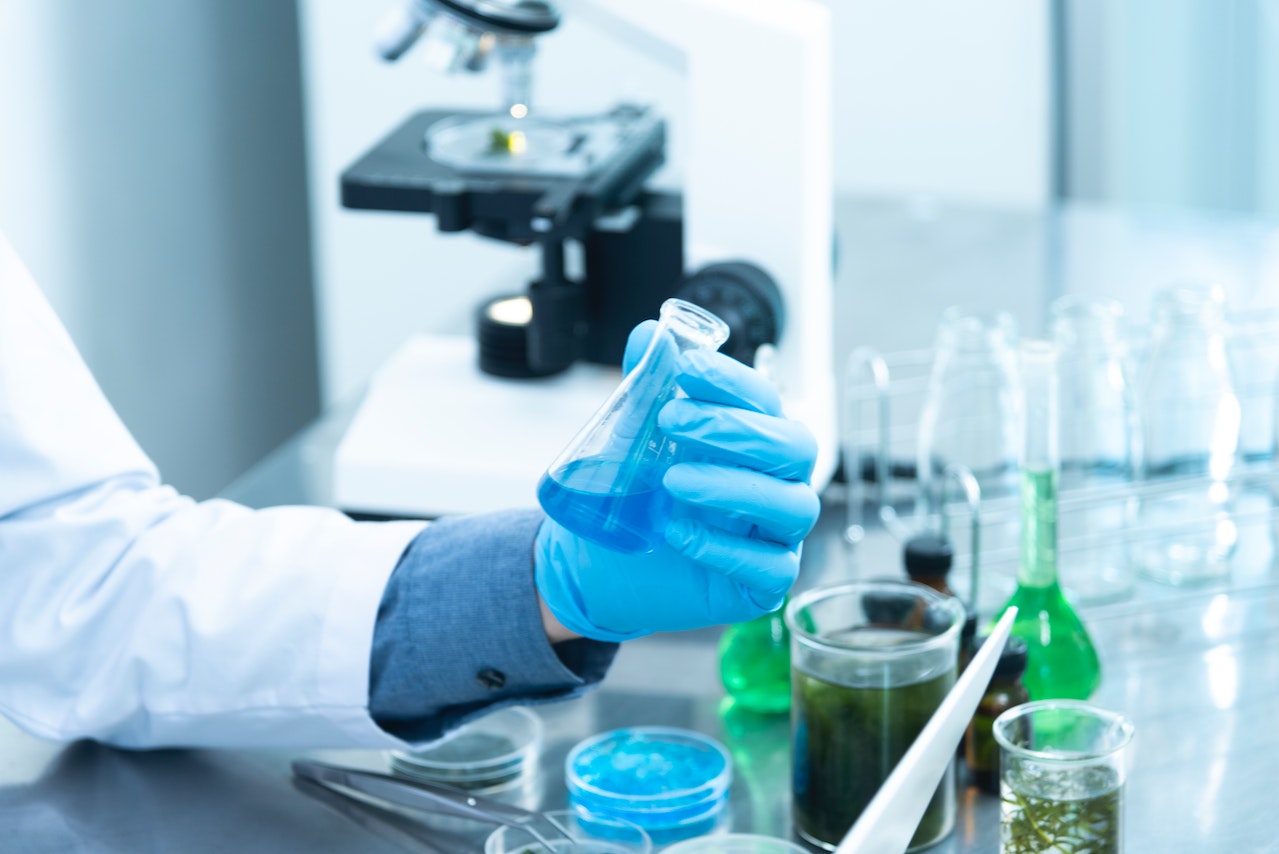
 How does a search engine work? Search engines are used when people ask questions and search for answers on the Internet. Search engine algorithms are
How does a search engine work? Search engines are used when people ask questions and search for answers on the Internet. Search engine algorithms are 

